Networks, Evolution, and the Question of Life
Streetlight 2017-10-18
It's a common misunderstanding among the lay public that individual genes, or rather particular sequences of DNA, simply 'code' for particular individual traits. The idea is that there is a one-to-one correspondence between gene and trait (in a slogan: "DNA makes RNA. RNA makes protein. Proteins make us"). For a variety of technical reasons, this is not quite the case. In general terms, the main reason is that the exact process of 'gene expression' (the process by which gene gives rise to trait) matters a great deal to the 'finished product', such that a single gene may in fact give rise to multiple outcomes, depending of the dynamics of the actual process of expression.
In fact, biologists these days often no longer even speak of individual genes, but of gene networks: it is the interaction of multiple genes, as well as with the specific developmental environment in which their expression takes place, that gives rise to particular traits. So note the change - no longer: one gene = one trait, but multiple genes (in interaction with the cellular environment) = multiple traits. One way to understand this is to look to Conrad Waddington's picture of the 'epigenetic landscape', as depicted below. The idea is that there are multiple 'ropes' which underlie the expression of any one trait, and cutting one, or even rearranging the organization of the ropes, may or may not affect the final outcome. Only by understanding the network and the interactions among it, may we understand how gene expression takes place:
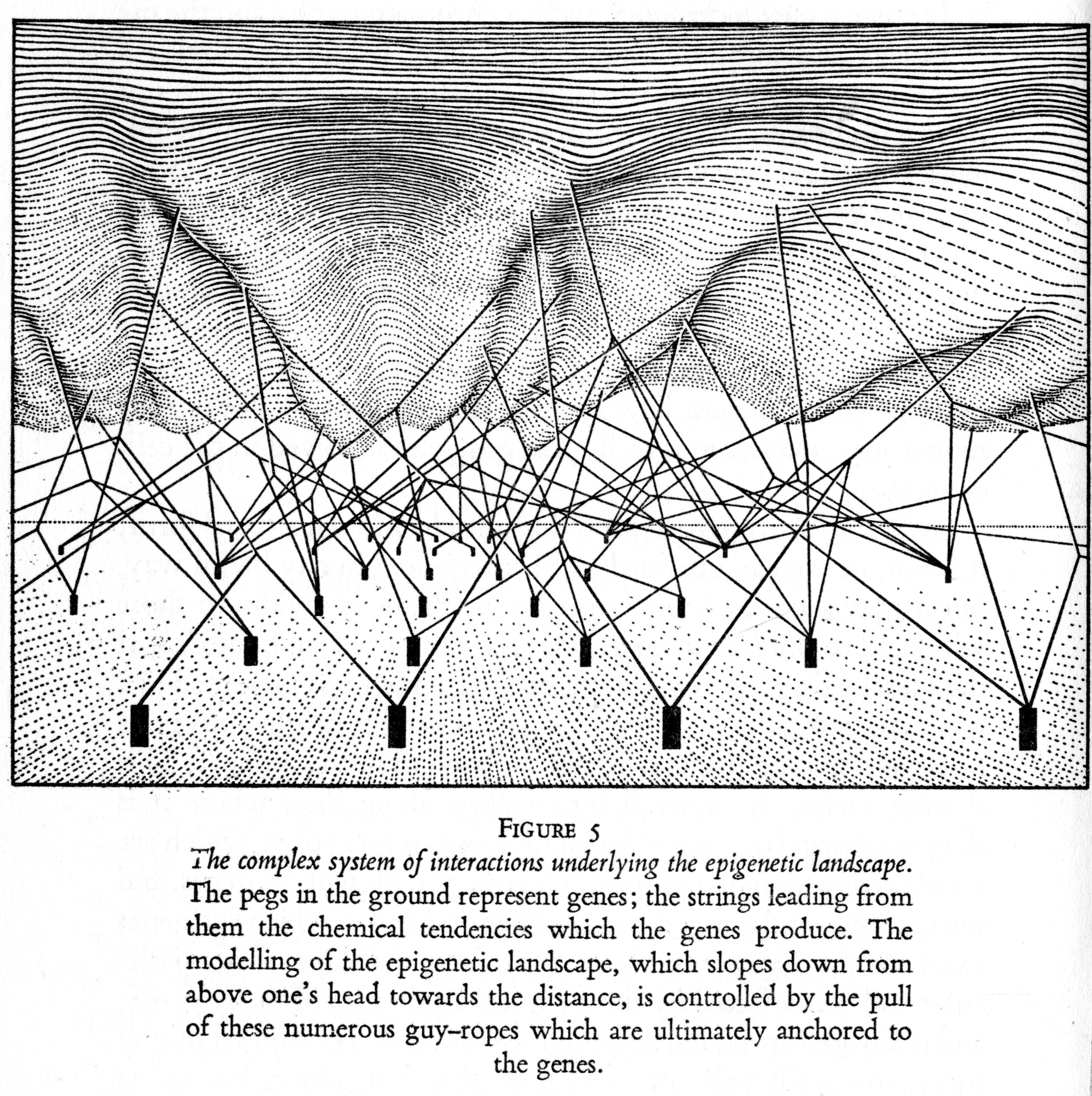
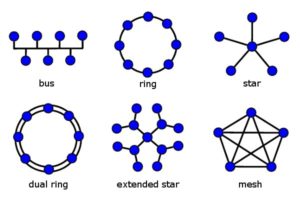
In terms of evolution, this matters because understood as such, the 'unit' of evolutionary variation can no longer be thought to be single genes, but rather, an entire genomic network (one consequence of this is that one can no longer talk about DNA as a simple 'blueprint' for life. Anytime you hear about a 'gene for X' or a 'gene for Y', you are most likely being sold simplified rubbish). Now, 'network' here is not a lightly used word: networks, after all, have very specific types of properties, and they are not biologically specific. For instance, networks have certain topologies or forms of organization (see picture below), along with other properties like robustness and plasticity, as well as threshold values and so on. What these are aren't that important right now, but note that applying network analysis to genome networks has proven to be quite scientifically fruitful: in terms of evolution, changing these network 'values' or properties is what leads to different phenotpyic outcomes.
Network diagram:

Network topologies:
Now, what's philosophically interesting to me about all this is that, if I understand the implications correctly, it throws into question the specificity of life itself, or rather what does and does not count as 'alive'. That is, if we think in terms of networks, how is it possible to think the specificity of life itself, insofar as the dynamics of genome networks are defined as much by extra-biological factors as they are biological ones? Because extra-biological factors are as just as important as biological factors in the process of gene expression, it becomes very hard to draw any kind of hard diving line between the two. This also follows, as a matter of principle, from the fact that networks are simply indifferent to the 'content' of the nodes which constitute them: it's all just a matter of the organization and threshold levels.
There's alot more to say, but as usual, I'm going to stop before I go on too long.
In fact, biologists these days often no longer even speak of individual genes, but of gene networks: it is the interaction of multiple genes, as well as with the specific developmental environment in which their expression takes place, that gives rise to particular traits. So note the change - no longer: one gene = one trait, but multiple genes (in interaction with the cellular environment) = multiple traits. One way to understand this is to look to Conrad Waddington's picture of the 'epigenetic landscape', as depicted below. The idea is that there are multiple 'ropes' which underlie the expression of any one trait, and cutting one, or even rearranging the organization of the ropes, may or may not affect the final outcome. Only by understanding the network and the interactions among it, may we understand how gene expression takes place:

In terms of evolution, this matters because understood as such, the 'unit' of evolutionary variation can no longer be thought to be single genes, but rather, an entire genomic network (one consequence of this is that one can no longer talk about DNA as a simple 'blueprint' for life. Anytime you hear about a 'gene for X' or a 'gene for Y', you are most likely being sold simplified rubbish). Now, 'network' here is not a lightly used word: networks, after all, have very specific types of properties, and they are not biologically specific. For instance, networks have certain topologies or forms of organization (see picture below), along with other properties like robustness and plasticity, as well as threshold values and so on. What these are aren't that important right now, but note that applying network analysis to genome networks has proven to be quite scientifically fruitful: in terms of evolution, changing these network 'values' or properties is what leads to different phenotpyic outcomes.
Network diagram:

Network topologies:

Now, what's philosophically interesting to me about all this is that, if I understand the implications correctly, it throws into question the specificity of life itself, or rather what does and does not count as 'alive'. That is, if we think in terms of networks, how is it possible to think the specificity of life itself, insofar as the dynamics of genome networks are defined as much by extra-biological factors as they are biological ones? Because extra-biological factors are as just as important as biological factors in the process of gene expression, it becomes very hard to draw any kind of hard diving line between the two. This also follows, as a matter of principle, from the fact that networks are simply indifferent to the 'content' of the nodes which constitute them: it's all just a matter of the organization and threshold levels.
There's alot more to say, but as usual, I'm going to stop before I go on too long.

Comments (75)
Really interesting. You, Apokrisis, fdrake, and a few others are often discourteous enough to try to tie the simplifications we play with in philosophy to the irreducibly complex real world.
This is not exactly the same thing, but what you've written reminded me of something I read in an essay by Stephen J. Gould. He wrote that the most important genes in terms of impact on the organism are often those that control the rate or sequence of expression of other genes. That made a lot of sense to me - that it is a network of interactions that you can't just hold in your mind as one piece as opposed to a stacking of bricks in a wall.
Also, can you describe the "extra-biological factors" you're talking about.
Have you not heard of biosemiosis yet? I am sure @apokrisis can explain this. I am not even being facetious here, this seems like a great example of how this fits into the biosemiosis framework.
Come now! Philosophy is no more or less rich than the 'real world' of which it speaks - you give it too little credit!
Quoting T Clark
Yeah, Gould was among the greatest champions of those who were dissatisfied with the 'gene-centrism' of biology, but even then, he hewed closely to the idea that genes were nonetheless the only units of heredity relevant to organisms, when this is more and more no longer thought to be the case.
The network approach really kicks sociobiology in the balls, which I'm sure Gould would have loved.
Where do ‘species’ fit in? Surely they rate a mention at least as ‘nodes’ in the network?
Doesn't it have to do with something like negentropy? Life appears to go against the Second Law of Thermodynamics, but it really isn't because its an open system that takes in free energy from its surroundings and produces heat and entropy back. Life is what is a negentropic open system.
I guess that may be more at the biophysics level of definition. But, are you asking whether life needs to have some sort of material substance like genetic material, to be considered life or is it more about the arrangement of the material? And if it is the arrangement of the material, what makes it different from any other arrangement of material? I'm just wondering if you can break down your post into a succinct question as far as the question of life you are proposing.
The problem I see here is you seem to be arguing that network topologies force their organisation on biological structure in a rather crystalline fashion. So something brute about possible topological arrangements would impose itself on biology and greatly restrict the variety of body structure that genes could encode.
But the argument is that genes are organised like neural networks and so embody a plastic intelligence. As you say, the genetic information would be distributed as a pattern of network weights. Yet in being a neural network, the network topology wouldn’t simply impose its own crystalline structure on the biology. Instead, the network could represent any kind of biological design as the “image it holds in mind”.
So just as a neural network can be trained to recognise any pattern, gene expression would be the same in reverse - the capacity to output any learnt pattern. The same intelligence of course helps to explain how the genome can recognise the current state of the body so as to respond appropriately in the first place. So really the genome is a pattern matching system.
And as such, there doesn’t seem any strong reason to suggest that the genome would be limited in the variety of states it could choose to represent. The limits such as they are would be more the logical ones - like the rules to generate bilateral symmetry and the repetition of segments - than topological ones. And that in itself is evidence that the gene networks must be operating at an intelligent level - actually relying on logic-like operations to switch and direct body development.
The OP is mostly concerned with developmental dynamics rather than evolutionary dynamics (despite the thread title!), so at this point the 'scale' of the discussion is limited to individual ontogeny rather than species-level phylogeny. There is, of course, the stupidly interesting question of former influences the latter, but that's probably a bit outside the bounds of the OP itself. I will say though, that one can think of a species as itself an individual (that is, subject to processes of individuation), which can itself populate a node among a larger ecological network.
The problem is that negentropy defines any type of organization, from whirlpools to star systems. That is, negentropy isn't specific to life, even as life counts as among the most magnificent examples of negentropic organization.
Quoting schopenhauer1
I guess the basic issue I'm grappling with is this: a genomic network is essentially indiscriminate with respect to what it counts as among its nodes - from a network perspective, whether a node is a biological element (say, DNA) or not is more or less irrelevant. All a network 'sees' is relations, structure, and threshold values. Of course, genes are necessary to, well, gene expression. But on the other hand, genes are also entirely inert, and they would in fact do and be nothing without the extra-biological scaffolding which enables that expression to take place in the first place. Consider this expanded, mirrored version of Waddington's epigenetic landscape:
Here it's more clear that life takes place 'in between' gene and environment, or the biological and the non-biological, which makes its exact specification very hard to place. Perhaps I can put it this way: if one can't identify the biological alone with life (insofar as there can be no such thing, strictly speaking, as 'biology alone'), then how exactly are we to situate 'life?'. I don't think, by the way, that this question can be answered categorically. I ultimately think that this is a political and even ethical question, rather than a strictly empirical one, but I want to specify, on the side of the empirical, as it were, why this would be the case.
Further, networks are individuated by more then their topological properties - you can have two networks with graph isomorphism: this states, roughly, that the relevance structure of the system is the same but have both different nodes (rendering the graph specific and individuated from its topological equivalents precisely because they are indexed to different genes) and also different flows on the network (more than one way to 'pass a current of information' through the graph). So it can be said that the topological properties of networks constrain the types of flow that pass through it, but the level of structural isomorphism is inappropriate for the analysis of the expression of particular genes or sets thereof.
The specificity comes in the application of general principles (like generating co-expression networks of specific genes) to individual gene clusters - discovering the clusters the process -. Specificity is always-already part of the developmental process for a given organism, but not necessarily part of the methodology of its analysis.
The problem I have is with the idea of "biology alone". Biology is not a self-contained set of substrates, but a fluid dynamic between certain macromolecules and the environment in a series of physical processes- some described probabilistically, some perhaps more straightforward (and all of it it perhaps biosemiotically). Thus it isn't just DNA, but the networks that they produce to create more complex processes. The networks that you describe may be analogous, if we were to isolate it in a network mapping way, but it is its situatedness, along with other biological substrates like DNA, cells, proteins and generally all the macromolecules that are found in lifeforms, that make it biological. The evolutionary history of how these networks came about and its unique way of solving problems using its structural constituents to influence its growth and development is what matters here.
Boy, this gets complicated. I can see - half genes from mom, half from dad. Trait A from mom, Trait B from dad. But if it's a network, that's 1/2 mom's network and 1/2 dad's network. Not even that. How do you inherit 1/2 a network? Why would it fit with the 1/2 network from the other parent? I'm not arguing against what you're saying.
Can a network change without a change in any specific gene? What establishes the network? Other genes? That would mean the network establishes the network. What mutates? I guess this type of genetic network would make the whole structure more stable, since everything's tied to everything else. As a civil engineer I say that the reason wood framed houses hold up so well is that there are thousands of nails and hundreds of pieces of wood all tied together.
Crap - you've given me more stuff to add to my reading list. I'm not as fast a reader as you guys.
If you cut one, or rearrange the organization of a set, and it doesn't affect the outcome (the expression of that trait), then it would be safe to say that that particular gene that was removed, or that particular arrangement of sets of genes that was rearranged, doesn't affect the expression of that trait. That is to say that there isn't a causal connection between that gene or arrangement of genes and that particular trait.
Biology is beholden to the laws of physics. You can't have the findings of two different fields contradict each other. Biology is just a sub-discipline of physics. Humans tend to put things into little boxes, including fields of science, which is ultimately an explanation of the world as a whole.
Quoting Harry Hindu
Really? The types of question appropriate in biology are a lot different from the types in physics. There are differences on the entities concerned, the relevant explanatory frameworks and the types of experiment conducted. That a biological system does not contradict any physical laws is one of the least interesting features of a biological system: the interesting ones concern its biology.
A reduction of biology to physics methodologically is pointless, philosophically a reduction of the living to the non-living is interesting though. Perhaps it's useful to say that the living is composed by the non-living in some manner, however.
'it's all made of particles!'
'is the tendency of negatively charged particles to repulse negatively charged particles made of particles?'
'no, it's something the particles do'
It seems to me that two different domains of investigation that ask different questions and end up NOT contradicting each other is what is really interesting.
Quoting fdrakeAgain, I don't see the distinction you are making. If a reduction of biology to physics is pointless, then so to is reducing the living to the non-living. Our bodies and natural selection must obey the laws of physics. If it didn't then we could all fly without the proper anatomy.
Quoting fdrake
Which is to say that the particle has a particular feature or property (it's negativity) that makes it do what it does. You seem to be implying that negativity could exist independent of particles. If it could, then why did you use the term, "particle" at all?
That a biological system doesn't contradict the laws of physics doesn't tell you much about the biological system. As you pointed out, stuff like air-resistance and aerodynamics enables things to fly; but doesn't make an incentive for things to fly. I meant the reduction of biology as a field of study to physics as a field of study is pointless - they're concerned about different things and use different methods. The ontological reduction from the living to the non-living might still be informative though.
I didn't intend to imply that the interaction of charges could exist independent of particles (requires charged particles and carrier particles). Just that it's something particles do, not something that they are. This was meant to highlight that the ontological reduction of stuff to particles doesn't even help much in describing what particles do (other than as an enabling condition for the study of particle behaviour); as an analogy to the reduction of biology as a field of study to physics as a field of study.
I'd rather keep the discussion related to what 'the reduction of the living to the non-living' suggests, rather than attempting to subordinate fields of study to each other. Just because I think the subordination of fields of study to each other is a different type of question, and is largely irrelevant, to answering and posing questions about life and the specificity of living organisms.
But just here is where the ambiguity over what then defines the boundary between life and not-life also enters into play, insofar as the topology between inside and outside (and this has nothing to do with the network topology we are talking about) becomes impossible to define (again, in the abstract). All you have, once again, is a developmental network largely indifferent to any boundaries that might be imposed form the top-down, as it were. Of course at this point it's tempting to do the scientific thing and simply follow the contours of the network itself: what does and doesn't make a difference to it in such and such cases? But then you get limit-cases like viruses which don't have the translation machinery for their DNA or RNA and need to hijack that of its host cells, or even - at the opposite end of the spectrum - life-support tech to sustain the body would would otherwise be unable to survive 'by itself'.
But because there's no a priori way to specify the limits of the biological here (one can only 'follow the system'), there's also no a priori way to rule out or in what should or shouldn't belong to any particular network. Life becomes not a problem of finding the right definition, but of finding the right 'application'. The question isn't "it this alive?', but 'it is fair to count this as an individual to which what we call life would or would not apply in the first place'? This is where one leaves behind the empirical and enters the realm of the ethical and the political. Evelyn Fox Keller gets at some of what I'm trying to bring out when she asks: "These terms are themselves ambiguous: what exactly is a gene, and what does it do? Even more troublesome is the ambiguity of the term environment. Do we mean it to refer to everything other than DNA, to the milieu in which the fertilized ovum develops, or to the factors beyond the organism that affect its development? Finally, there is also the question, contributions to what?" (Fox-Keller, The Mirage of a Space Between Nature and Nurture).
I agree! I spoke of 'biology alone' precisely in order to render the notion a bit ridiculous; that was the point of the expanded Waddington diagram, to show that it is impossible to speak, in any coherent way, of biology alone with respect to life. And you're also right that this means that what matters is the evolutionary history of any particular network, such that we can't say what ought to or not belong to any particular network in advance (isn't one of the marvels of evolution it's ability to hijack or incorporate the environment into it's dynamics?). But this, I want to say, has conceptual consequences for what we understand 'life' to be, and how fragile a notion it is.
May I introduce you to genetic redundency and genetic robustness, or more specifically canalisation, if we're talking about genes alone. They're dear friends, be nice to them.
One super interesting thing to bring up in relation to this - I might start another thread on this down the line - is in following Robert Rosen's contention that biology is, contrary to what is commonly thought, a more general science than physics, insofar as biological systems have a richer repertoire of causal entailments than do physical ones. Physical systems thus being a more limited field of study, even if they qualitatively make up more of the universe. This is one of those lovely thoughts, I think, that spurred me to study biology in some depth - want to know physics? Study biology! :D
I remember how much you hate possible world semantics for modal logic, but I think some idea of possibility/necessity modality is useful here to sharpen the claim that a demarcation criterion between the biotic and abiotic is impossible - at least as it concerns the generating processes of organisms.
I assume that a demarcation criterion is largely an epistemological device, and nature would not care about the distinction between the divided factors apart from differences in generalised processes that influence them.
Imagine at some future time there was a complete list of all the things which could influence the expression of an arbitrary assemblage of genes. Call this collection P. At any time before this collection is made, there will be a subset Q(t) of P that represents the current list of all influences on genetic expressions.
Considering the structure of Q(t), there are likely to be subclasses that are united by shared properties - generalities in genomic networks indexed to sets of co-expressing genes. Can we tell at any time whether the set of properties is exhaustive, and that we have provided a spanning partition of P generated by the properties? If the set of studied gene expressions were fixed and finite, in principle this would be possible. Not in practice however, there is an absurd combinatorial explosion whenever you're dealing with genome subunits, nevermind sets of genome subunits paired with genome subunits...
One way to represent P would be the set of all common expression properties, represented by their network bearers. In order to decide that we have all the properties that constitute P we would need a demarcation between factors that influence the development of genes and those that do not. Specifically, we would need to be able to infer from some particular Q(t) that there are no more possible types of gene expression influencing entities or processes - that we truly have a spanning partition of P, even if we do not know all its elements.
Evolution conditions Q(t) and the generated property list approximating P, since there will always be novel environmental scenarios. In particular, this ensures that novel categorisations are always indexed with a set of environmental parameters, at least insofar as the development of developmental processes is concerned. The indexing role the environment would play in this kind of study would ensure that environmental conditions are in some sense dense - ubiquitously represented and always very 'close' in the sense of accompaniment and foundation to accounts of changes in developmental biology- within the study of developmental processes as they relate to their environments. However, it is also likely that the environment is conceived as that set of things 'outwith' the 'biological' constituents of the developmental process. Despite the environment's ubiquity (denseness) within the analytic concepts of the development of developmental biology. This operationally equates the concepts of environmentality and exogeny. (You can observe the converse happening in ecology btw, the equation of an ecology with its interiority to description)
Whether the demarcation problem for the biotic and the abiotic is decideable within or using information from these theories would then depend on the extent to which the abiotic is operationally defined as the biologically exogenous, and whether it is operationally necessary to treat it as such.
Responding with quote from a physicist friend: "I'm uncomfortable with science that deals with anything that isn't a state variable. (temperature, pressure)"
I don't see incentive as part of the equation. Things behave in certain ways as a result of how they were designed. There was no incentive prior to, or the cause of, flight. Flight occurred as a result of natural selection acting on genetic mutations over eons. By saying there is an incentive is projecting your own purposes onto reality, as if reality has reasons, or incentives, to design things. It doesn't. "Design" isn't even an appropriate term to use to describe what natural selection does, as there is no incentive, purpose, reason, or goal that natural selection has prior to the process itself taking place.
Quoting fdrake
Then if it wasnt implied that negativity was a separate feature of a particle, then it must be that it is an integral feature of that particle. Repelling other particles is what that particle does as a result of its negativity. If it didn't repel other particles then we couldn't say that possessed negativity.
Everything that something does is limited by and determined by its shape and the laws of physics. Incentives are still limited by the laws of physics.
That's what I meant. Teleological language is useful to paraphrase stuff like that.
It's relative, not absolute purpose. For a phenotypic variation to be differentially replicated,* it must provide some survival or reproductive advantage relative to other members of the population. Surviving and reproducing is not an absolute purpose, but it's how the game is played.
I remember seeing an episode of NOVA or something where they were testing chickens as part of a theory that wing flapping might evolve as a sort of turbo boost for running-- that is, helps you survive which helps you reproduce.
* That sounds like it could be a TMBG song about evolution.
Yes. The interesting point about genomic networks is that their internal processing structure could be - in principle - uncrackable and forever hidden. Can we reconstruct the way a neural network executes its function even with full knowledge of the weights of its nodes?
If the functionality is multirealisable, then a knowledge of some particular state of task-adapted componentry does not give a simple theory of the functional dynamics of the network.
We could still hope to model genomics at a higher level. That’s why I’m thinking of a description in terms of general logical principles. Like the repetition of units (as in segmented body plans) or timing information when it comes to regulating tissue growth and developmental symmetry breakings or bifurcations.
That is a good point. A big constraint on the whole gene story is that there is this strong selection for the whole damn business being evolvable too. The genome both has to regulate development and also do its best to expose individual traits to the winnowing force of selection.
So while a distributed genome behaves as a processing network, it still wants to offer up traits on an individualised basis, not the whole package.
This was the argument for why we go from the simple gene rings of bacteria to the chromosomes of multicellular life. The chromosomes, with their recombination shuffling of the gene deck, look to be a mechanism that both preserves the coherence of the developmental network and allows traits to be exposed to evolution in a suitably individualised fashion.
The evolution of sex - the separation of the immortal germ-line - is another important step. It means the genome that runs the everyday body is kept out of the evolutionary fray. The trait-level view is the specialist role played by gametes, or sperm and egg.
I'm not a biologist, and have only tertiary knowledge in developmental biology.
For a specific gene network, precise estimation of parameters would be the goal. To think of a genomic network as structurally isomorphic to a neural network is probably possible, but it will remove both specificities. I doubt, though I could be wrong, that genomic networks are necessarily concerned with message passing in continuous or quasi-continuous time like neural networks are; and seem to be constructed through a correlational analysis of genes to traits and genes to genes in terms of expression. I know there are counterexamples when analysing cellular differentiation.
I think this is a truism. But it will not diminish the analysis of a specific genomic network.
Dynamical systems theory is already being used. Street's reference to canalisation has a link to bifurcation theory.
Edit: I would rather attempt to keep this on topic than to sidetrack the discussion into your semiotic metaphysics, though.
Physics certainly constrains biology. But what then isn't constrained is by definition free to happen. And this freedom is what demands further modelling.
In fact, this freedom is something we have begun to generalise by talking about computation, information, negentropy, modelling relations, semiosis, etc.
So rather than biology just being some small and accidental expression of something physics failed to forbid, we are realising that it leads to a whole realm of "mind". The fact that physics creates a hard baseline of constraints in itself opens up a matchingly definite counter-possibility of a hard superstructure of rather absolute freedoms. Information becomes a real source of causality acting back on the world in evolutionary fashion.
That is why talk of genenomic networks being like neural networks in having a "separate adaptive intelligence" makes sense. A test of how much this is the case would be to consider the resilience that genomes show if you try to knock out one developmental pathway, and yet the genome can reorganise to still produce the desired functional outcome.
Quoting StreetlightX
Yep.
That would be the research question. We do have a variety of neural network architectures as candidates. The genome could be just like of those, or something different.
Quoting fdrake
Surely genomic networks would have to be very much always monitoring the state of the cell? The biophysics now makes the point that cellular machinery is always on the point of breaking down and so needing the right nudge to build itself back up.
Direct evidence that timing matters is how the most time-critical regulation - that of the respiratory chains of mitochondria - is handled right on the spot by mitochondrial genes.
So most of the mitochondria genes have migrated to the central chromosomes. The time lags are not so critical. But where fast response is needed, the genes are kept right where the reactions are taking place.
Quoting fdrake
Of course. But this is now different if instead of the dynamics being something directly physical - the self-organising dynamics at a "chemical" level - we are talking about a dynamics-based information model.
So this is about the genome as analog computation. A dynamical description might apply. But at the level of disembodied information rather than embodied physics.
It is just the same as the problem of talking about the representation of the world in the brain. Patterns of neural activity don't just simply "look like" the physics of the world they are modelling. There is certainly some topographical relationships preserved, but also a hierarchy of functional specialisation. Activity in one small blob of the visual cortex produces our sense of colour, another that of motion.
So the old mental picture of the genome was a "flat" network at best. It was just some kind of straight transcription layer with not particular internal complexity.
Once we start talking about neural networks, we are asking just how much internal hierarchical organisation, and hence "mind-like" complexity, there is going on. A very different ball game.
The nodes in neural networks are placed there by modellers and use a message passing algorithm to update parameters linking the nodes. The nodes in gene expression networks are discovered through a kind of cluster analysis. The nodes mean different things, the nodes are generated by different things. Nevertheless if there are flows on the networks there will be general mathematical descriptions of the flows. The flows will mean very different things for the different systems.
Generally, just a great big citation needed on the material in your post.
I see a confusion here. Human-made machinery of course is engineered to be physics-free and so the hardware is completely generic in terms of its informational capacity. The network boils down to a collection of binary switches that represent a memory state. So there is a strong division between memory and processing. The divorce between the setting of weights and the dynamics of any "processing interaction with the world" is rather absolute.
From that mechanical basis, a neural network will then evolve some operational state. That can then be described in functional terms. Some collection of nodes, with their pattern of weights, will "act like a particular feature detector". We will want to point to that functional element of the internal model as a cluster that analyses. But really, that seems a heuristic rather than a properly motivated claim. A mathematically soft one, rather than a rigorously grounded one.
Then with real-life genomic networks, we can't even really claim that they even want to start with actual digital circuitry - the binary switches that enforce a strong divorce between memory states and dynamical behaviour. Genes may approach some kind of digitalism, and yet still be essentially "analog all the way down".
For me, this is still the great unsolved riddle of life and mind. Sure, mostly science presumes a digital ground - life and mind are just super-complicated machinery. But ontologically, I take the semiotic view where creativity, spontaneity and uncertainty are irreducible (even if pragmatically constrainable). And so confusion will arise if genomic networks are still imagined as having to rest on mechanically stable foundations - actual little hardware switches.
Quoting fdrake
Hah. Yep. I'm afraid I've deliberately steered clear of gene expression for the past 15 years, rather waiting for the science to sort itself out. But biophysics has suddenly got interesting, so gene regulation again becomes interesting for me as now it is easier to see what it actually has to control. And that has become much less due to the fact that molecular machinery does most of the hands-on regulation.
A cell isn't a bag of chemistry but itself a highly regulated, almost machine-like, network of entropy flow or dissipative structure.
Again, the hierarchy of control is becoming apparent. Molecular machinery is like a whole new level of regulation in itself, whereas before there was only a genetic blueprint and a chemical soup.
The following may be somewhat tangential to your concerns, but what the hell?
In the ordinary sense we all broadly understand the distinction between "living' and 'non-living'. As per Wittgenstein; there is no determinate essence that could define the difference; rather the difference is found in the whole context of networks of "family resemblances" between phenomenal manifestations that we think of divergently as either living or non-living.
Since I have been reading his works lately Michel Henry is also much in my thoughts at the moment. He draws what he sees as an all-important distinction between Life and Being. Since Being is the always-already externalized world, life, for Henry, cannot be found there at all, but we will find only entities which do or do not manifest phenomena that we might understand to be 'life functions'.
So what would you say are some consequences of this idea that what matters in biology is the situatedness of the molecules and the environment in its unique historical evolutionary development?
Language isn't teleological. It's that most of human minds are, and they project that onto reality in how they use a language. There are others that try to avoid that, and in so doing, recognize the faults of the language and attempt to create new terms to use that doesn't lead to one simply paraphrasing, but actually getting at what it is that we are talking about independent of any subjective projections.
That isn't freedom. Computers are designed to perform many different functions - from creating a document and printing it, to surfing the internet, to playing 3D games. This wide range of things that computers do doesn't mean that they are free to do something they weren't designed to do, like fly off your desk and make you breakfast.
Hm, there's a bit of an ambiguity here, I think, already at the level of formulation: any genomic network is already a space of possibilities, such that some parts of the network may be active in any particular process of expression, while other parts may not be - and just which parts are and are not may be dependent on certain (regulatory) genetic and epigenetic conditions. This means, further, that what even counts as 'a' network is not fixed, and individuation is itself dependent on the parameters of any one investigation - what we count as belonging or not belonging to 'a' network (the space of possibilities), and by extention, what we count as 'a' network to begin with, is itself not something fixed in advance. Of course this is just the scientific process: fix the boundaries of the phenomena you want to study, hold all else equal, then poke around. When you ask then:
- The problem, it seems to me, is that even if we could get around the combinatoric issues, any exhaustive list of properties would be in some sense only so by fiat. And if so, I'm not sure how much we can milk the distinction between P and Q(t) to really speak about any demarcation between the biotic and the abiotic.
The other, intimately related, conceptual issue I see is that because genomic networks are complex, the activation or deactivation of certain parts of the network (via regulation) may alter the very possibility space itself: what was once an 'influence' which would never have been able to play a role in the expression of a certain trait, becomes an influence, or vice versa. And this change may have knock-on effects with respect to other 'possible influences' as well; things get confusing, I think, because at stake are second-order possibilities: 'possible possibilities', as it were. And again, at this point, I'm not sure how stable any distinction between P and it's subset Q(t) might be...
Sure, sure, but the question is what kind of licence developmental biology allows us when it comes to speaking about life in the terms it provides. The context here is quite rigoursly defined, and is not at odds with Witty's insights.
Quoting Janus
Yeah, I'm familiar with Henry - I think he's an absolute genius - but I also think his conception of 'Life' qua self-affecting, pathic 'Subjectivity' is - there's no nice way to put this - entirely vacuous. I wrote a long, multipost thread in the old forum about this - on his conception of immanence in particular - which is lost to time now, but yeah, I've never found his mobilization of the concept of 'Life' very - if at all - useful, unfortunately.
Quoting apokrisis
Think. Can a Universal Turing computer fly off the desk and make you breakfast?
How does this issue have implications for our thoughts about "what does and does not count as alive"?
It's not news that organisms like us need oxygen to stay alive. That gives us no reason to speak as though oxygen is alive.
The fact that the boundary between an organism and its environment is fuzzy and permeable doesn't mean there's no difference between the organism and its environment, and doesn't mean there's no difference between animate and inanimate objects.
Of course, developmental biology will add to our stock of "phenomenal manifestations that we think of divergently as either living or non-living" and may even change our minds as to whether we think of particular phenomena as living or non-living. It may even lead some thinkers to dissolve the distinction altogether which is what you and fdrake seem to have been alluding to in some of your comments.
Quoting StreetlightX
It doesn't surprise me that you would think that given the presuppositions that are inherent in your approach. In a way I agree; life, as conceived by Henry is vacuous from the viewpoint of the determinate world of "ek-static phenomena". However, it is anything but vacuous in terms of living affection; which is really Henry's point.
It's not so much the goal to 'dissolve' the distinction as to 'denaturalize' it, to make it an object less of scientific analysis than political and ethical judgement.
Quoting Janus
But Life is not vacuous from the ek-static point of view: as Henry is everywhere at pains to point out, Life is the quite literally the essence of manifestation; Henry's critique of the whole phenomenological tradition before him is that time and time again, it comes across Life, only to end up ignoring it. The problem, if anything, is the other way around - it's the Henry's own philosophy is almost entirely uncritical, in the Kantian sense - Life is the essence of manifestation, but it cannot, ironically, account for the manifestation of the ek-static phenomena which it everywhere underlies. This is not an original critique, and has been pointed out time and time again by many commentators of Henry's work - and entirely rightly, I think.
Here is Renaud Barbaras: "Although he discovers auto-affection at the heart of aIl givenness at a distance, Henry never heads down the opposite path to discover how auto-affection leads into intentionality, how we can go from immanence to transcendence.... Henry cannot provide answers to these questions precisely because he argues that they concern two completely impenetrable regimes of appearance. " (Barbaras, The Essence of Life: Desire or Drive?); and Dan Zahavi: "Henry operates with the notion of an absolutely self-sufficient, non-ekstatic, irrelational self-manifestation, but he never presents us with a convincing explanation of how a subjectivity essentially characterized by such a complete self-presence can simultaneously be in possession of an inner temporal articulation; how it can simultaneously be directed intentionally toward something different from itself; how it can be capable of recognizing other subjects (being acquainted with subjectivity as it is through a completely unique self-presence); how it can be in possession of a bodily exteriority; and finally how it can give rise to the self-division found in reflection:" (Zahavi, Subjectivity and Immanence in Michel Henry). See also Ray Brassier's devastating reading of Henry in his Alien Theory, which I won't quote here.
Finally, your own point that Life isn't vacuous form the point of view of Life is... well, just a tautology. But again, this is all very off-topic!
Because the topology between 'inside' and 'outside' at stake here is different: it's not just that there are 'organisms' on the one side and 'oxygen' on the other; it's that epigenetic and environmental influences are already 'on the side' of life, or rather the organism, to begin with. That's the whole point of focusing on networks: whether the nodes in a genomnic network are biotic or abiotic is a matter of sheer indifference from the point of view of the network, which can only 'see' relations, topologies, and threshold values. While it's true, as others have pointed out, there is a kind of specificity provided by the spatio-temporal dynamics of the cellular environment itself, this only serves, as I've argued, to worsen the ambiguity because those dynamics themselves also cannot be neatly parsed along biotic/abiotic lines.
The problem is that life traverses both 'sides' in the manner of a mobius strip or klein bottle, where the distinction between inside and outside cannot really, be made:
I still don't see what difference you're suggesting.
So far it seems perhaps you're mistaking an abstract mathematical representation of some interactions in a physical system, for the physical system itself.
It's a matter of indifference, "from the point of view" of a geometrical abstraction, what the abstraction is an abstraction of. It's no surprise if a geometrical representation of complex physical interactions omits a great deal of information, and no surprise if distinctions that are relevant to us in some contexts are not relevant in the framework of that geometrical representation.
The organism ends at its own fuzzy boundary, but the environment does not. Organisms are interwoven with their environments, and are entirely composed of material from their environments over time. The material we're made of is only borrowed for a while.
That material is organized into a physically coherent structure in space and time, so the organism is distinguishable in its environment according to principles of biological organization; but it remains in constant physical interaction with a local physical context, in a continuous exchange of matter and energy.
Given a particular "set of genes", which may be characterized physically; and a range of possible environmental factors, which can be characterized as distinct sets of physical factors; there corresponds a range of phenotypical outcomes, which can be characterized as distinct sets of physical states or physically determined capacities or "traits" of the organism. The organism, its genes, its environment, and its traits all have concrete physical existence in space and time, and can be distinguished accordingly.
Now you say some mathematician comes along to represent those concrete physical relations abstractly by sketching beautiful geometrical topographies in his notebook. And in that abstract mathematical representation, perhaps, "it is a matter of indifference" whether the nodes are biotic or abiotic.
Does that mean anything more than that, once you abstract from real physical context, and represent complex physical features of an environment as mathematical "nodes", you may lose sight of the complex physical features, and forget to look back and forth between the enchanting sketch and the real world it is designed to represent?
Depending on what sort of representation we're discussing, I suppose it may also be a matter of indifference whether the organisms and traits and environmental factors represented in the diagram are actual or only possible -- such maps may represent relations of sets of possible genetic codes, possible environmental factors, possible traits -- and not a single real thing in the world. In fact there's a difference between my genes, my traits, and the environments I've actually passed through over time, on the one hand, and all the environments I could possibly have passed through in a mathematician's dream, and all the traits I might have had if things had gone otherwise, on the other hand.
That may be "a matter of indifference from the point of view of the representation", too. But it's not a matter of indifference to real organisms that suffer real environments.
OK, but that leaves me wondering what criteria the political and ethical judgements will ideally be based upon.
Quoting StreetlightX
You seem to have misunderstood what I was alluding to. I wasn't trying to suggest that Henry wants to stress that life is vacuous "from the ek-static point of view". Of course every phenomenon is a manifestation of life for Henry. The point is that life cannot be analyzed from the ek-static point of view; an intentional account of it cannot be given, and no account of life can be inter-subjectively corroborated. It is on account of this that I said life (I probably should have that the idea of life is vacuous instead, because obviously life, however you conceive it cannot be vacuous in itself) is vacuous from the perspective of "the truth of the world". and I think this is pretty much behind what you are saying when you say that
Quoting StreetlightX
Thanks for the other references, anyway, but, really I am just beginning to explore Henry's ideas, and I would much rather flounder my way to a creative misreading, if that's what it is to come to; than be 'corrected' by 'expert' secondary opinions.
Ah fair enough, that makes more sense.
Quoting Janus
Heh, that's fine too, and despite my critical take on Henry, I still think he's an absolutely incredible philosopher who deserves to be widely studied and read. I think his intitutions regarding intentionality are absolutely right, even if I think his 'solution' is entirely wrong! Glad you're reading him nonetheless.
Quoting Janus
Man, that's a whole other kettle of fish, but, at a minimum, I'd say that part of the challenge of both ethics and politics - the myriad differences between them notwithstanding - is that there can be no ideal criteria, only criteria immanent to the context(s) of judgement in which judgement must be exercised. But again, that's outside the scope fo this thread.
But I'm talking about a process: the process of gene expression, and the question of how this process, which necessarily traverses both biotic and abiotic elements, entails an inability to situate life clearly on the side of the biotic. If anything, the abstraction lies in breaking down the process into it's analytic elements and ignoring its holistic aspects. If, on the other hand, I speak about the process in terms of a network, it is because network thinking best brings out the processual nature of what is at stake. It is no use, as such, in simply speaking of individual entities like 'organism', 'environment', etc - none of these capture of processual specificity of what is at stake here.
I think perhaps this is a mountain out of a molehill. While there are some processes that are abiotic mixed in with the biotic (like the networks with biotic molecules and environment), this does not mean that we cannot distinguish abiotic and biotic processes altogether. Biotic processes are ones that have a mix BUT have the known constituent parts that comprise biological molecules. Thus if ATP is going, DNA sequencing, etc. etc. then more than likely this is biological. As we discussed biology is not just the networks abstracted but instantiated in the very evolutionary way it has "solved" the problems of the environment. This is something that has not been replicated and can perhaps be mapped, but mapping it in abstraction is not the same thing.
But this would include say, ecosystems and river catchments. In any case I'm not arguing that we can't distinguish between biotic and abiotic processes: my point is rather that life itself traverses both such that life cannot be defined in strictly biotic terms. And to be extra clear, I'm also not arguing that we can't distinguish between life and not-life, only that such a distinction cannot be 'read off' the phenomena themselves in any straightforward way, if only because what exactly would and would not count as 'a' phenomena is precisely what is in question: a question of individuation.
Again though, biological systems are a series of networks that are hierarchical. They are relational, and if some important components of the "nodes" are taken away, the system stops functioning. However, the core constituents and their evolutionary development are what matters here to distinguish it from any old system.
This description would apply to literally any complex system, living or not. And besides, to repeat for the third time, the question is not whether or not we have a criteria for a living system or not, but whether one can discern whether or not such a criteria would apply to begin with. If you keep ignoring the fact that at issue is a question of individuation (what does and does not count as 'a' system), you'll miss what I'm trying to say.
I don't think so because you are forgetting the part about biological constituents with the unique evolutionary ways that the organism uses to solve problems in the environment.
Quoting StreetlightX
I think though these two problems are related in the case of your problem. Where are the limits of biological systems? Should there be limits to any system which is open and sharing some form of information? Do you accept that there can be discrete units that relate with other discrete units? As i said there is a hierarchy for each discrete unit of organism.
But this is just circular then: biological systems are systems with biological constituents...
Quoting schopenhauer1
I don't understand what you're asking with these questions. Could you be more specific?
I don't think so, do other systems reproduce, metabolize, using the same unique set of tools (biological molecular parts)? Do other systems evolve in the unique way biological systems do? No and no, and thus we start making distinctions between this system versus other systems. But I know, this is not what you thread is supposed to be about.
Quoting StreetlightX
It sounds like you're asking things like "What makes an ecosystem different than an organism?" and I wanted to see what your thoughts were about what makes an organism an organism.
But this is the first time you've mentioned metabolism or reproduction. You previously spoke simply of hierarchical networked systems, that were, in a way not yet specified, biological.You seem to be filling in your definition as you go along. But - and this is about the fourth time I'm making this point, and I'm afraid I will not make it again - whether or not you do have an adequate definition of the living is not in question. It is a question of application, which is why I spoke about limit cases like viruses or life-support systems with fdrake, and which even Harry mentioned. Further, I never once asked 'what makes an ecosystem different than an organism', because I have been quite clear that I've been speaking about processes, and not 'units' of things. You seem to keep going over ground that is not relevant to what is being addressed. Perhaps I have not been clear, but it is tiring trying to correct for viewpoints I do not hold.
Quoting StreetlightX
Perhaps I can put it this - not super precise - way: the question is not 'what is alive?', but 'what is alive?'. This latter is the question of individuation, of what counts-as a-life, a question which I think is opened by a reflection on the process of gene expression.
Quoting StreetlightX
I think the first quote there makes it difficult for some of us to understand what approach is suggested by the second quote. You seem to rule out exactly what most of us would be doing if we set out either to choose a criterion for life or apply it. It makes it look as though you want to ask, not whether something that eats is living, but whether eating is living, and that can't be what you're saying.
Is the connection that an organism is "composed" of such processes?
The analogy that came to my mind was the relation between phonemes and morphemes. Morphemes, as the smallest units of semantic content, are by definition made of parts that do not have semantic content. As such, it's no use looking to the phonemes to determine whether a given object is a morpheme; there is in a sense nothing special, nothing directly related to semantics about phonemes. You can't read off morpheme-hood. (You can do some things, given more rules: say what could be a morpheme in a given language, etc.)
But now I seem to be doing the sort of thing the first quote would discourage; and the second quote suggests that the issue is what a unit of semantic content could possibly be, given that the unit is composed of non-semantic units.
Am I at all close to understanding your point?
:)
This indifference, I think, opens up a fascinating speculative possibility: that life itself can be entirely divorced from the biotic. To pursue this thought in a different register, consider the following line of speculation from the evolutionary biologists Eva Jablonka and Marion Lamb, who envisage a possible future in which the DNA inheritance system is altogether replaced by higher-order one - and that, not only could this possibly happen, but that something like it might have already happened with respect to our already existing DNA system:
"In existing organisms, which all have a nucleic acid–based inheritance system, it is inconceivable that the DNA inheritance system will be eliminated by another one that operates at a higher level. But theoretically it is possible that one heredity system can replace another. It may well have happened at an early stage in the evolution of life, during the murky period between chemical and biological evolution. Many theorists suggest that heredity during these early stages was not based on nucleic acids, and that the nucleic acid systems came later and replaced the primitive heredity systems. Maybe such a replacement will also occur in the distant future— if we create intelligent, reproducing, and evolving robots, they may eventually eliminate us. This would be equivalent to the elimination of one heredity system by another”. (Jablonka and Lamb, Evolution in Four Dimensions)
J+L are here not talking of course, about gene expression but inheritance systems in evolution more generally, but the principle is the same: that life itself may be considered as something entirely separate from the biotic. Life would thus be a formal principle rather than a material one: so long as the process and its organisation are kept in place, the exact ‘instantiation’ of the formal principle is - from a very specific perspective - a matter of sheer indifference (I take inspiration also from Robert Rosen and Nicholas Rashevsky’s imperative to "throw away the matter and keep the underlying organisation”, when speaking about biological systems).
The final step here is to see that formal systems, by definition, do not provide an index of their own applicability. That is, they simply say: ‘anything that meets these organisational criteria belong to the class of systems so named living’. But what does and does not count as ‘a’ thing here is not something that can be read off the phenomena itself: someone on a life support system might be said to be structurally coupled to that system, without which they would die: but the individuation of what counts and does not count here is a matter of judgement, not a matter of empirical study - hence the moral dilemmas we face when, in some circumstances, we decide on turning off a life support system or removing an breathing tube. Again, I’ve gone on too long and there’s still lots to say. But hope this fills out some possible gaps.
Got it. I believe I am following you, and sometime tomorrow I will move on to thinking about it.
I think you need to remove the ambiguity from your categories of "process" to understand the criticism which has been directed at your approach.
First, we understand in general, "processes". Then we have the more particular "living processes". It would appear like we would have some principles, whereby we distinguish living processes, as a particular type of process. Classically one would refer to a living being as carrying out the process, and this would determine a living process.
You have removed this living thing, which is assumed to carry out the living process, so from my perspective you have no means for distinguishing living processes from other processes. Now it makes no sense to classify a process as a living process because there is no basis for distinguishing a living process from a non-living process.
It appears like what you are trying to do, and this would be a real challenge, is to make living processes into the more general category. In this case, all processes would be distinguished in relation to being alive. Sub-categories, different types of processes, would all be related to each other through what you call what counts as alive. Here I can see three categories, was alive, is alive, and will be alive.
Don`t see anything to pick at here. So long as the problem space for the demarcation of life and not-life is the space of possible genomic networks, rather than specific ones. Not that this space of possibilities is independent of the considerations relevant to the networks.
I don`t understand why you would think it would be by fiat, since the construction, or discovery, of genomic networks is a fortiori the conceptual space for that demarcation at least as far as this thread is concerned. Rather, the inclusion of types of factors in them is the conceptual space. This isn`t a criticism of what you said, rather to elicit more information. Why at any given point would it be by fiat?
I don't think that this is a particularity of genomic networks, they only instantiate the problems that epigenetics rises for the distinction of life. Inclusion in a specific genomic network requires that a node is a gene, the more general picture you raise of the epigenetic landscape is the appropriate space of concepts for tracing the implications for the demarcation of biotic and abiotic factors.
I also think it's likely that abiotic and biotic factors are likely to have an operational definition within the study of genomic networks that doesn't coincide completely with the (supposedly) philosophical distinction between life and not-life.
The appropriate conceptual space for this argument is something like genomic networks as a problem instantiation of epigenetics for the demarcation between life and not-life, so decisions (subarguments) should be considering the scope of genomic networks and how their internal distinctions relate to life and not-life, and further how this relates to epigenetic effects - then how that counterfactual space of concepts (study of genomic networks -> epigenetic effects) relates to the demarcation problem for life.
It looks to me, though I may be misreading you, that you are committing something of a category error (at least if the above typology of the space of problems is accurate), confusing the inclusion within specific genomic networks and its formal undecideability with respect to the biotic and the abiotic with the general features of the study of genomic networks (which is where epigenetics comes in). This paper incorporates this distinction methodologically.
Edit: I see you addressed some of this in response to @Srap Tasmaner
Are there any known philosophies that DO think that biological constituents must be in the picture along with the process and its organization (i.e. contra Rosen perhaps)? If so, what is their reasoning? I'm just trying to kick off a dialectic to ensure you are not overlooking counter-examples.
What is there in this world that isn't a process, or in process, or the abstract result of some process?
The process of being alive involves biotic and abiotic materials: At any time in which a living organism exists, there are materials currently integrated as the organism, materials in the process of being integrated into the organism, materials in the process of being disintegrated from the organism, and materials that are not, are not being, and have not been integrated with the organism. The boundary of the organism's integrity is not the same as the generic boundary of biotic and abiotic organization: There's a great deal of recycling of material from each organism to others.
How does this sort of account "ignore the holistic aspects" of the relation between organism and environment, or the holistic aspects of the process of living?
It's still not clear to me why you seem to think everyone's been in the dark about the porousness of the relation between organism and environment. It's still not clear to me how abstract representation of features of the relation of genetic code, environment, and traits in terms of mathematical networks and topologies is supposed to be a remedy for the alleged misconception.
Of course life is a process. Of course the organism changes over time in constant interaction and exchange with the rest of its environment. Recent investigation into the way "traits" result from various combinations of genetic material and environmental factors is a welcome refinement of our view of that interaction. Perhaps it's especially valuable as a correction of a too-narrow misconception of the relation between genetic code and traits. But the idea that external factors -- like the quantity and quality of food, water, sunlight, air, and company -- influence the path of an organism's development seems a very old idea indeed.
Note, just so there's no confusion - this is a response to a discussion that took place 6 months ago. It was one of my favorites. Reading on the web, I came across the linked article which I think is interesting and relevant.
https://www.quantamagazine.org/how-many-genes-do-cells-need-maybe-almost-all-of-them-20180419/